Meta.Morf 2012 – A Matter of Feeling
A MATTER OF FEELING –CURATORIAL STATEMENT.By ALEX ADRIAANSENS
A Matter of Feeling. By Alex Adriaansens, director V2_, Rotterdam. Advertising has made us chasing cars and clothes, we have working jobs we hate so we can buy shit we don’t need. Our great depression is our lives. We’ve all been raised by television to believe that one day we’d all be millionaires, and movie gods, and rock stars, but we won’t. We’re slowly learning that fact. And we’re very, very pissed off. Tyler Durden in “The Fight Club” The social welfare state is falling…
LaunchA MATTER OF FEELING.by RACHEL ARMSTRONG
A Matter of Feeling. Introductional essay for the Meta.Morf 2012 conference. The cosmos is composed of many different species of stardust and despite our advanced, secular knowledge we imagine these primordial substances give rise to a universe, fashioned in our own image. Meta.Morf is a reflection on a new kind image, which is evolving in a diverse set of arts practices at the start of the twenty-first century. Intriguingly, its portrait of our universe is far more autonomous and sensitive than the one that has…
LaunchALTERNATE EMBODIMENTS / PROSTHETIC HEAD.By STELARC
ALTERNATE EMBODIMENTS / PROSTHETIC HEAD We are living in an age of excess and indifference. Of prosthetic augmentation and extended operational systems. There is now a proliferation of biocompatible components in both substance and scale that allows technology to be attached and implanted into the body. A turbine heart has been engineered that is more robust and reliable than previous artificial hearts. It circulates the blood continuously without pulsing. In the near future you may rest your head on your loved ones chest. He is…
LaunchFEELING MATTER: EMPATHY AND AFFINITY IN THE HYLOZOIC SERIES.By Philip Beesley.
Feeling Matter: Empathy and Affinity in the Hylozoic Series. Our work on the Hylozoic Series of interactive sculptures attempts to offer an anatomy of subtle boundaries that expands the physiology of individual human bodies. Might it be possible to suspend judgement about what is ‘me’ and what is clearly not? Rather than weakness, the ambivalence implied by such a suspension might be an enabling quality. The Hylozoic Series attempts to expand the space that lies between clearly defined personal territories, transforming it into a felt…
LaunchFICTIONAL INFLUENCES.By NEIL SPILLER
Fictional Influences. I do not make any distinction between fiction and reality. I do not believe in reality – I see it as a capitalist fabrication as much fiction as any fiction. I am a second order cyberneticist who believes that we “make” our individual, personal worlds by operating and “building” within them. Whatever that “building” is, whether it is a conversation, poetry, prose, the design of objects or buildings. We are changed by conversations and the relationships we make with what we perceive as…
LaunchMIND TIME MACHINE AS A LIVING TECHNOLOGY.By Takashi Ikegami
Mind Time Machine as a Living Technology. Any basic science can lead to innovative applications. Artificial life (alife) studies is no exception. The purpose of living technology is to bring to fruition the concepts developed through the study of artificial life, such as self-reproduction, autonomy, enaction, robustness, open-ended evolution, evolvability, and so on, in a real-world context (Ikegami, 2009). By increasing our understanding of how we can connect artificial systems with natural environments, we can further our development of a theoretical framework that situates artificial…
LaunchMONOLAKE [DE NL]Sep. 28 @ BYSCENEN
Byscenen Friday September 28 Monolake Ghosts in Surround presents the material of the Ghosts album as a full surround sound experience plus real time generative video. The visual side of the Ghosts in Surround tour is a contribution by the Dutch audiovisual artist Tarik Barri [NL] with whom Robert Henke [DE] is collaborating since early 2009. “…Precisely crafted earthshaking beats, rough dirty noises, deep bass, wide lush soundscapes and little sonic creatures inhabiting a fascinating planet in which a lot of things go badly wrong and nothing is taken for granted –…
LaunchOPEN REEL ENSEMBLE [JP]Sep. 28 & 29 @ BYSCENEN
Byscenen Friday 28th and Saturday 29th of September The return of magnetic tape ! OPEN REEL ENSEMBLE hides an unusual group of 5 young musicians led by Ei Wada (who recruited 4 friends from university for his project) and its stand-out feature is its signature instrument: old rolls of magnetic tape that the group’s members connect to computers from the latest generation, creating unprecedented melodies, tones and sounds and providing a live show that defies any possible comparison with anything else you have ever seen on a…
LaunchFROST [NO] Sep. 29 @ BYSCENEN
CONCERT @ Byscenen Saturday September 29! FROST Live: Aggie Peterson, Per Martinsen. Visuals: Petra Hermanova. Frost is the musical project of creative couple Aggie Peterson and Per Martinsen. They’ve released two albums together (“Melodica” in 2001 and “Love! Revolution!” in 2006) and a dozen singles and remixes. They have toured all from Paris to London, and played several big festivals like Denmark’s Roskilde and Norway’s Øya Festival. Between them they’ve also been involved in a variation of other projects, such as staging art shows for…
LaunchHERE TO GO:Art, Counter-Culture and the EsotericSymposium Oct. 20 @ NOVA
Forum Nidrosiae Symposium Saturday October 20 @ NOVA. Featuring CARL ABRAHAMSSON, ANDREW M. McKENZIE, GARY LACHMAN, KENDELL GEERS, JESPER AAGAARD PETERSEN and KAREN NIKGOL. Curated and moderated by MARTIN PALMER. I talk a new language. You will understand. – Brion Gysin All lectures will be in English. Throughout the ages, artists have worked with ways of expressing the “inexpressible”, and with bridging the gap between the material world and our inner space. There is an intimate relationship between art and the spiritual. Through the 20th…
LaunchRALF BAECKER [DE]
Irrational Computing (2011) IRRATIONAL COMPUTING investigates material, aethetics and potential of digital processes. The basic raw materials of our surrounding information technology are semiconductor crystals such as silicon, quartz or silicon carbide, which, thanks to today’s advanced microtechnology and extremely sophisticated procedures, are processed into transistors or integrated circuits (IC), with the materiality of modern microprocessors having long since ceased to be graspable. The extreme miniaturization and the black-box set-up elude visual interpretation. The Installations circuit runs counter to the developments in information technology, representing the…
LaunchPHILIP BEESLEY [CA]
Epiphyte Grove – Hylozoic series (2011 – 2012) The Hylozoic Ground experimental architecture series developed by architect Philip Beesley has been expanded and refined by researchers, engineers and designers from around the world. It is an immersive, interactive environment that moves and breathes around its viewers, creating an environment that can ‘feel’ and ‘care’. Next-generation artificial intelligence, synthetic biology, and interactive technology create an environment that is nearly alive. This new technology has, according to the London Times, “the power to be the dominant aesthetic of…
LaunchØYVIND BRANDTSEGG [NO]
Installation for a pedestrian bridge (2012) Co-produced by Meta.Morf / TEKS – Trondheim Elektroniske Kunstsenter. The installation is based on rhythm and sound generated by pedestrians passing the Shipyard Bridge (Flower Bridge, Trondheim). 8 contact speakers and 8 contact microphones are mounted to the underside of the bridge walkway. Through exploring the rhythmic expression, the installation relates to development and negotiation of meaning in language. Rhythm patterns recorded from the audience walking across the bridge are recorded and used as source material for the development…
LaunchWIM DELVOYE [BE]@ TSSK
CLOACA N˚ 5 (2006) 2006 / 390 x 70 x 330 cm / Mixed Media Wim Delvoye is a Belgian neo-conceptual artist known for his inventive and often shocking projects. Much of his work is focused on the body. “Delvoye is involved in a way of making art that reorients our understanding of how beauty can be created.” Delvoye is perhaps best known for his digestive machine, “Cloaca”, which he unveiled at the Museum voor Hedendaagse Kunst, Antwerp, after eight years of consultation with experts in…
LaunchDRIESSENS & VERSTAPPEN [NL]
Breed / E-volved Cultures (2001 – 2011) Breed Breed is a computer program that uses artificial evolution to grow very detailed sculptures. The purpose of each growth is to generate by cell division from a single cell a detailed form that can be materialised. On the basis of selection and mutation a code is gradually developed that best fulfils this “fitness” criterion and thus yields a workable form. The designs were initially made in plywood. Currently the objects can be made in nylon and instainless steel…
LaunchXANDRA VAN DER EIJK [NL]
Momentum (2011) Momentum is a silent symphony, a dance between artist and installation, with the liquid (like time itself) flowing at its own pace. Xandra Van Der Eijk Momentum shows the process of decay and the human incapability to overcome it. It is a spacious, analog installation with a 4 to 6 -day cycle, guided by a very slow, continuous performance of assisting the installation in order for it to work, resulting in strong graphic, 4m long prints. Because of certain ingredients in the mixture,…
LaunchPETER FLEMMING [CA]
Instrumentation (2012) Piano strings, mechanical saltwater dimmer, plywood transducer table, electromagnetic pickups, stepper motor actuated light dimmer, hardware, electromagnetic coils, custom circuits, drums. Instrumentation is an electro-mechanical sound installation inspired by resonance. The gallery installation preserves a sense of the makeshift, having evolved from studio experiments, using a limited palette of tools and readily available materials. Spanning two separate but connected spaces, different aspects of the work are presented in each. Unlikely loudspeakers improvised from buckets, drums, salvaged windows and hand-wound electromagnetic coils occupy a…
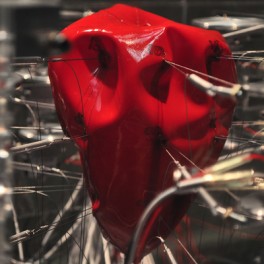
LaunchMARKUS KISON [DE]
Pulse (2012) Co-produced by Meta.Morf / TEKS – Trondheim Electronic Arts Centre & V2_, Rotterdam, 2012 Rubber, acrylic glass, motors, arduino microcontrollers, laptop, processing applications, Internet connection, Blogger.com API. Pulse is a live-visualization of recent emotional expressions, written on private weblog communities like wordpress.com. Weblog entries are compared to a list of emotions, which refers to Robert Plutchik’s seminal book Psychoevolutionary Theory of Emotion published in 1980. Plutchik describes eight basic human emotions in his book: joy, trust, fear, surprise, sadness, disgust, anger, and anticipation. He developed a diagram…
LaunchKIANOOSH MOTALLEBI [UK]
Terrestrialball (2010) A small spherical object made with the 94 naturally occurring elements on Earth. It measures 1 inch in diameter. Terrestrial Ball is a small, spherical object made of the 94 elements that occur naturally on earth. It is at once a tangible look at the world we live in, a memento of our home and an object that relates in an elementary way to every other object and substance that has ever been made. Meteors – bits of other worlds that fall onto…
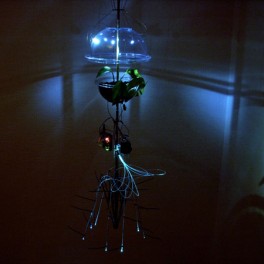
LaunchGUTO NOBREGA [BR]
Breathing (2009) Breathing is a small step towards new art forms in which subtle processes of organic and non-organic life may reveal invisible patterns that interconnect us. Breathing is a work of art based on a hybrid creature made of a living organism and an artificial system. The creature responds to its environment through movement, light and the noise of its mechanical parts. Breathing is the best way to interact with the creature. This work is the result of an investigation of plants as sensitive…
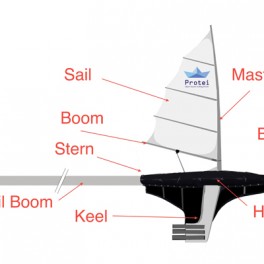
LaunchPROTEI / OIL COMPASS [FR US UK PL DE]
Shape-shifting Open Hardware Sailing Robot for Ocean Sensing and Cleaning PROTEI Protei is an international collaborative project born at the intersection of art, interaction design and science. Initiated by Cesar Harada in the Gulf of Mexico during the BP oil spill, Protei is developed primarily to intercept Oil Spills sheens drifitng down the wind with a long oil absorbant “tail” sailing up the wind. With its shape-shifiting hull, it appropriates existing technologies in an innovative low cost design that can be implemented on the short term to…
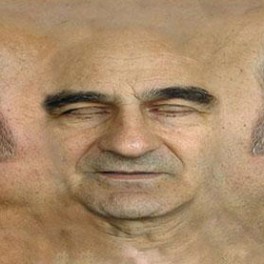
LaunchSTELARC [AU]
Prosthetic Head (2003) Prosthetic Head is an automated, animated and reasonably informed artificial head that speaks to the person who interrogates it. The PROSTHETIC HEAD project is a 3D avatar head, somewhat resembling the artist, that has real time lip-synching, speech synthesis and facial expressions. Head nods, head tilts and head turns as well as changing eye gaze contribute to the personality of the agent and the non-verbal cues it can provide. It is a conversational system which can be said to be only as…
LaunchZIMOUN [CH]
Woodworms, wood, microphones, sound system (2007 – 2012) ”Using simple and functional components, Zimoun builds architecturally-minded platforms of sound. Exploring mechanical rhythm and flow in prepared systems, his installations incorporate commonplace industrial objects. In an obsessive display of curiously collected material, these works articulate a tension between the orderly patterns of Modernism and the chaotic forces of life. In an obsessive display of curiously collected material, these works articulate a tension between the orderly patterns of Modernism and the chaotic forces of life. Carrying an emotional depth,…
LaunchSOUNDSPACE 1:1Public installationSep. 27 – Oct. 28@ Trondheim
Architecture students cooperate with students from the music technology program on the theme of Sound and Space in city spaces. To study architecture is largely about developing a consciousness about the meaning of space, of its possibilities and its limitations. The first phase of this learning process is to explore and create space – to get personal experiences with the spatial. The first assignment one gets as an architecture student at the Norwegian University of Science and Technology is a complete small-scale building project. The…
Launch